
Garrett R.H., Grisham C.M. - Biochemistry (1999)(2nd ed.)(en)
.pdf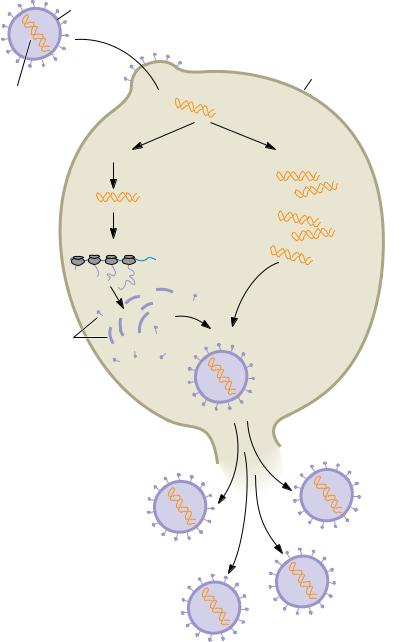
1.6 ● Viruses Are Supramolecular Assemblies Acting as Cell Parasites |
31 |
Often, viruses cause the lysis of the cells they infect. It is their cytolytic properties that are the basis of viral disease. In certain circumstances, the viral genetic elements may integrate into the host chromosome and become quiescent. Such a state is termed lysogeny. Typically, damage to the host cell activates the replicative capacities of the quiescent viral nucleic acid, leading to viral propagation and release. Some viruses are implicated in transforming cells into a cancerous state, that is, in converting their hosts to an unregulated state of cell division and proliferation. Because all viruses are heavily dependent on their host for the production of viral progeny, viruses must have arisen after cells were established in the course of evolution. Presumably, the first viruses were fragments of nucleic acid that developed the ability to replicate independently of the chromosome and then acquired the necessary genes enabling protection, autonomy, and transfer between cells.
Protein coat
Host cell
Entry of virus Genetic 
 genome into cell material
genome into cell material
(DNA or RNA)
Transcription |
Replication |
 RNA
RNA
Translation
Coat proteins |
Assembly |
|
FIGURE 1.25 ● The virus life cycle. Viruses are mobile bits of genetic information encapsulated in a protein coat. The genetic material can be either DNA or RNA. Once this genetic material gains entry to its host cell, it takes over the host machinery for macromolecular synthesis and subverts it to the synthesis of viral-specific nucleic acids and proteins. These virus components are then assembled into mature virus particles that are released from the cell. Often, this parasitic cycle of virus infection leads to cell death and disease.
Release from cell
32 Chapter 1 ● Chemistry Is the Logic of Biological Phenomena
PROBLEMS
1.The nutritional requirements of Escherichia coli cells are far simpler than those of humans, yet the macromolecules found in bacteria are about as complex as those of animals. Since bacteria can make all their essential biomolecules while subsisting on a simpler diet, do you think bacteria may have more biosynthetic capacity and hence more metabolic complexity than animals? Organize your thoughts on this question, pro and con, into a rational argument.
2.Without consulting chapter figures, sketch the characteristic prokaryotic and eukaryotic cell types and label their pertinent organelle and membrane systems.
3.Escherichia coli cells are about 2 m (microns) long and 0.8m in diameter.
a.How many E. coli cells laid end to end would fit across the diameter of a pin head? (Assume a pinhead diameter of 0.5 mm.)
b.What is the volume of an E. coli cell? (Assume it is a cylinder,
with the volume of a cylinder given by V r2h, where 3.14.)
c.What is the surface area of an E. coli cell? What is the surface- to-volume ratio of an E. coli cell?
d.Glucose, a major energy-yielding nutrient, is present in bacterial cells at a concentration of about 1 mM. How many glucose
molecules are contained in a typical E. coli cell? (Recall that Avogadro’s number 6.023 1023.)
e.A number of regulatory proteins are present in E. coli at only one or two molecules per cell. If we assume that an E. coli cell contains just one molecule of a particular protein, what is the molar concentration of this protein in the cell?
f.An E. coli cell contains about 15,000 ribosomes, which carry out protein synthesis. Assuming ribosomes are spherical and have a diameter of 20 nm (nanometers), what fraction of the E. coli cell volume is occupied by ribosomes?
g.The E. coli chromosome is a single DNA molecule whose mass is about 3 109 daltons. This macromolecule is actually a linear
array of nucleotide pairs. The average molecular weight of a nucleotide pair is 660, and each pair imparts 0.34 nm to the length of the DNA molecule. What is the total length of the E. coli chromosome? How does this length compare with the overall dimensions of an E. coli cell? How many nucleotide pairs does this DNA contain? The average E. coli protein is a linear chain of 360 amino acids. If three nucleotide pairs in a gene encode one amino acid in a protein, how many different proteins can the E. coli chromosome encode? (The answer to this question is a reasonable approximation of the maximum number of different kinds of proteins that can be expected in bacteria.)
4.Assume that mitochondria are cylinders 1.5 m in length and 0.6 m in diameter.
a.What is the volume of a single mitochondrion?
b.Oxaloacetate is an intermediate in the citric acid cycle, an important metabolic pathway localized in the mitochondria of eukaryotic cells. The concentration of oxaloacetate in mitochondria is about 0.03 M. How many molecules of oxaloacetate are in a single mitochondrion?
5.Assume that liver cells are cuboidal in shape, 20 m on a side.
a.How many liver cells laid end to end would fit across the diameter of a pin head? (Assume a pinhead diameter of 0.5 mm.)
b.What is the volume of a liver cell? (Assume it is a cube.)
c.What is the surface area of a liver cell? What is the surface-to- volume ratio of a liver cell? How does this compare to the surface- to-volume ratio of an E. coli cell (compare this answer to that of problem 3c)? What problems must cells with low surface-to- volume ratios confront that do not occur in cells with high surface-to-volume ratios?
d.A human liver cell contains two sets of 23 chromosomes, each set being roughly equivalent in information content. The total
mass of DNA contained in these 46 enormous DNA molecules is 4 1012 daltons. Since each nucleotide pair contributes 660 dal-
tons to the mass of DNA and 0.34 nm to the length of DNA, what is the total number of nucleotide pairs and the complete length of the DNA in a liver cell? How does this length compare with the overall dimensions of a liver cell? The maximal information in each set of liver cell chromosomes should be related to the number of nucleotide pairs in the chromosome set’s DNA. This number can be obtained by dividing the total number of nucleotide pairs calculated above by 2. What is this value? If this information is expressed in proteins that average 400 amino acids in length and three nucleotide pairs encode one amino acid in a protein, how many different kinds of proteins might a liver cell be able to produce? (In reality, liver cells express at most about 30,000 different proteins. Thus, a large discrepancy exists between the theoretical information content of DNA in liver cells and the amount of information actually expressed.)
6.Biomolecules interact with one another through molecular surfaces that are structurally complementary. How can various proteins interact with molecules as different as simple ions, hydrophobic lipids, polar but uncharged carbohydrates, and even nucleic acids?
7.What structural features allow biological polymers to be informational macromolecules? Is it possible for polysaccharides to be informational macromolecules?
8.Why is it important that weak forces, not strong forces, mediate biomolecular recognition?
9.Why does the central role of weak forces in biomolecular interactions restrict living systems to a narrow range of environmental conditions?
10.Describe what is meant by the phrase “cells are steady-state systems.”
FURTHER READING
Alberts, B., Bray, D., Lewis, J., et al., 1989. Molecular Biology of the Cell, 2nd ed. New York: Garland Press.
Goodsell, D. S., 1991. Inside a living cell. Trends in Biochemical Sciences 16:203–206.
Koonin, E. V., et al., 1996. Sequencing and analysis of bacterial genomes. Current Biology 6:404–416.
Lloyd, C., ed., 1986. Cell organization. Trends in Biochemical Sciences 11:437–485.
Loewy, A. G., Siekevitz, P., Menninger, J. R., Gallant, J. A. N., 1991. Cell Structure and Function. Philadelphia: Saunders College Publishing.
Pace, N. R., 1996. New perspective on the natural microbial world: Molecular microbial ecology. ASM News 62:463–470.
Further Reading |
33 |
Service, R. F., 1997. Microbiologists explore life’s rich, hidden kingdoms. Science 275:1740–1742.
Solomon, E. P., Berg, L. R., Martin, D. W., and Villee, C., 1999. Biology, 5th ed. Philadelphia: Saunders College Publishing.
Wald, G., 1964. The origins of life. Proceedings of the National Academy of Science, U.S.A. 52:595–611.
Watson, J. D., Hopkins, N.H., Roberts, J. W., et al., 1987. Molecular Biology of the Gene, 4th ed. Menlo Park, CA: Benjamin/Cummings Publishing Co.
Woese, C. R., 1996. Phylogenetic trees: Whither microbiology? Current Biology 6:1060–1063.

“If there is magic on this planet, it is contained in water.”
LOREN EISLEY
Inscribed on the wall of the National
Aquarium in Baltimore, MD
OUTLINE
2.1 ● Properties of Water
2.2 ● pH
2.3 ● Buffers
2.4 ● Water’s Unique Role in the Fitness of the Environment
34
Chapter 2
Water, pH, and Ionic
Equilibria
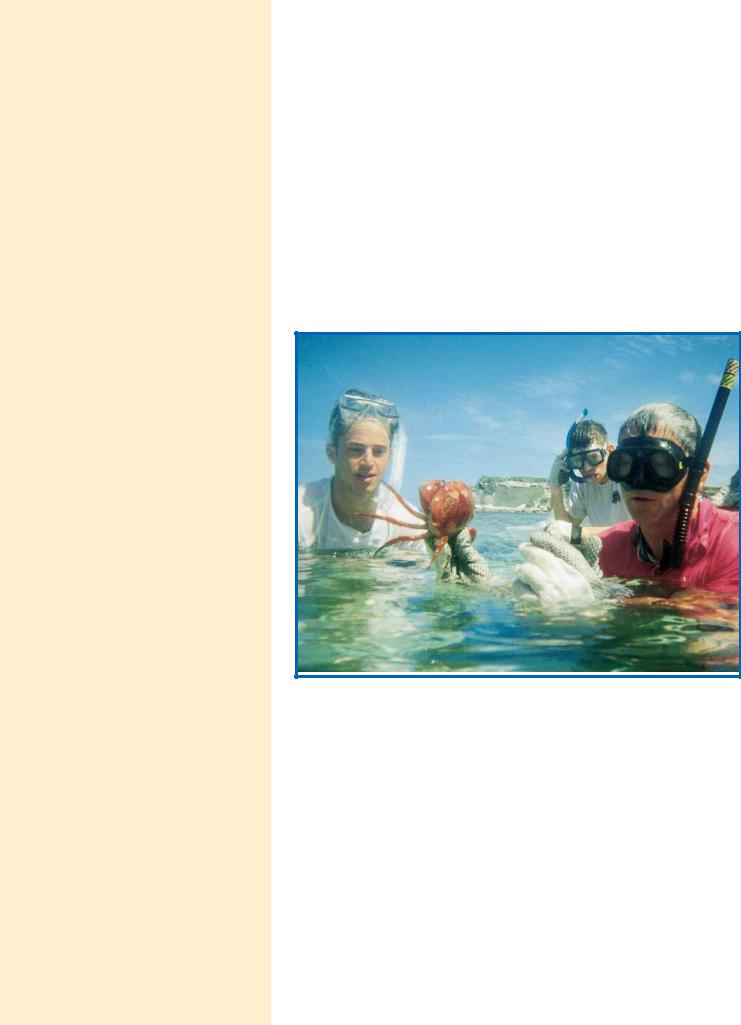
Some of the magic: Students and teacher view a coral crab in Graham’s Harbour, San Salvador Island, the Bahamas. (Lara Call)
Water is a major chemical component of the earth’s surface. It is indispensable to life. Indeed, it is the only liquid that most organisms ever encounter. We alternately take it for granted because of its ubiquity and bland nature or marvel at its many unusual and fascinating properties. At the center of this fascination is the role of water as the medium of life. Life originated, evolved, and thrives in the seas. Organisms invaded and occupied terrestrial and aerial niches, but none gained true independence from water. Typically, organisms are constituted of 70 to 90% water. Indeed, normal metabolic activity can occur only when cells are at least 65% H2O. This dependency of life on water is not a simple matter, but it can be grasped through a consideration of the unusual chemical and physical properties of H2O. Subsequent chapters establish that water and its ionization products, hydrogen ions and hydroxide ions, are crit-

ical determinants of the structure and function of proteins, nucleic acids, and membranes. In yet another essential role, water is an indirect participant—a difference in the concentration of hydrogen ions on opposite sides of a membrane represents an energized condition essential to biological mechanisms of energy transformation. First, let’s review the remarkable properties of water.
2.1 ● Properties of Water
Unusual Properties
In comparison with chemical compounds of similar atomic organization and molecular size, water displays unexpected properties. For example, compare water, the hydride of oxygen, with hydrides of oxygen’s nearest neighbors in the periodic table, namely, ammonia (NH3) and hydrogen fluoride (HF), or with the hydride of its nearest congener, sulfur (H2S). Water has a substantially higher boiling point, melting point, heat of vaporization, and surface tension. Indeed, all of these physical properties are anomalously high for a substance of this molecular weight that is neither metallic nor ionic. These properties suggest that intermolecular forces of attraction between H2O molecules are high. Thus, the internal cohesion of this substance is high. Furthermore, water has an unusually high dielectric constant, its maximum density is found in the liquid (not the solid) state, and it has a negative volume of melting (that is, the solid form, ice, occupies more space than does the liquid form, water). It is truly remarkable that so many eccentric properties should occur together in a single substance. As chemists, we expect to find an explanation for these apparent anomalies in the structure of water. The key to its intermolecular attractions must lie in its atomic constitution. Indeed, the unrivaled ability to form hydrogen bonds is the crucial fact to understanding its properties.
Structure of Water
The two hydrogen atoms of water are linked covalently to oxygen, each sharing an electron pair, to give a nonlinear arrangement (Figure 2.1). This “bent” structure of the H2O molecule is of enormous significance to its properties. If H2O were linear, it would be a nonpolar substance. In the bent configuration, however, the electronegative O atom and the two H atoms form a dipole that renders the molecule distinctly polar. Furthermore, this structure is ideally suited to H-bond formation. Water can serve as both an H donor and an H acceptor in H-bond formation. The potential to form four H bonds per water molecule is the source of the strong intermolecular attractions that endow this substance with its anomalously high boiling point, melting point, heat of vaporization, and surface tension. In ordinary ice, the common crystalline form of water, each H2O molecule has four nearest neighbors to which it is hydrogen bonded: each H atom donates an H bond to the O of a neighbor, while the O atom serves as an H-bond acceptor from H atoms bound to two different water molecules (Figure 2.2). A local tetrahedral symmetry results.
Hydrogen bonding in water is cooperative. That is, an H-bonded water molecule serving as an acceptor is a better H-bond donor than an unbonded molecule (and an H2O molecule serving as an H-bond donor becomes a better H-bond acceptor). Thus, participation in H bonding by H2O molecules is a phenomenon of mutual reinforcement. The H bonds between neighboring molecules are weak (23 kJ/mol each) relative to the HOO covalent bonds (420 kJ/mol). As a consequence, the hydrogen atoms are situated asymmetrically
2.1 ● Properties of Water |
35 |
FIGURE 2.1 ● The structure of water. Two lobes of negative charge formed by the lonepair electrons of the oxygen atom lie above and below the plane of the diagram. This electron density contributes substantially to the large dipole moment and polarizability of the water molecule. The dipole moment of water corresponds to the OOH bonds having 33% ionic character. Note that the HOOOH angle is 104.3°, not 109°, the angular value found in molecules with tetrahedral symmetry, such as CH4. Many of the important properties of water derive from this angular value, such as the decreased density of its crystalline state, ice. (The dipole moment in this figure points in the direction from negative to positive, the convention used by physicists and physical chemists; organic chemists draw it pointing in the opposite direction.)
36 Chapter 2 ● Water, pH, and Ionic Equilibria
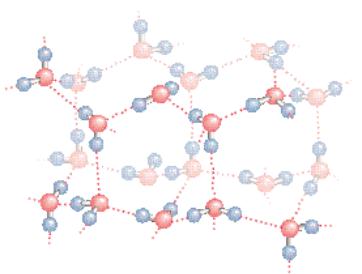
● The structure of normal ice. The hydrogen bonds in ice form a threedimensional network. The smallest number of H2O molecules in any closed circuit of H-bonded molecules is six, so that this structure bears the name hexagonal ice. Covalent bonds are represented as solid lines, whereas hydrogen bonds are shown as dashed lines. The directional preference of H bonds leads to a rather open lattice structure for crystalline water and, consequently, a low density for the solid state. The distance between neighboring oxygen atoms linked by a hydrogen bond is 0.274 nm. Because the covalent HOO bond is 0.995 nm, the HOO hydrogen bond length in ice is 0.18 nm.
between the two oxygen atoms along the OOO axis. There is never any ambiguity about which O atom the H atom is chemically bound to, nor to which O it is H-bonded.
Structure of Ice
In ice, the hydrogen bonds form a space-filling, three-dimensional network. These bonds are directional and straight; that is, the H atom lies on a direct line between the two O atoms. This linearity and directionality mean that the resultant H bonds are strong. In addition, the directional preference of the H bonds leads to an open lattice structure. For example, if the water molecules are approximated as rigid spheres centered at the positions of the O atoms in the lattice, then the observed density of ice is actually only 57% of that expected for a tightly packed arrangement of such spheres. The H bonds in ice hold the water molecules apart. Melting involves breaking some of the H bonds that maintain the crystal structure of ice so that the molecules of water (now liquid) can actually pack closer together. Thus, the density of ice is slightly less than the density of water. Ice floats, a property of great importance to aquatic organisms in cold climates.
In liquid water, the rigidity of ice is replaced by fluidity, and the crystalline periodicity of ice gives way to spatial homogeneity. The H2O molecules in liquid water form a random, H-bonded network with each molecule having an average of 4.4 close neighbors situated within a center-to-center distance of 0.284 nm (2.84 Å). At least half of the hydrogen bonds have nonideal orientations (that is, they are not perfectly straight); consequently, liquid H2O lacks the regular latticelike structure of ice. The space about an O atom is not defined by the presence of four hydrogens, but can be occupied by other water mole-
cules randomly oriented so that the local environment, over time, is essentially uniform. Nevertheless, the heat of melting for ice is but a small fraction (13%) of the heat of sublimation for ice (the energy needed to go from the solid to the vapor state). This fact indicates that the majority of H bonds between H2O molecules survive the transition from solid to liquid. At 10°C, 2.3 H bonds per H2O molecule remain, and the tetrahedral bond order persists even though substantial disorder is now present.
Molecular Interactions in Liquid Water
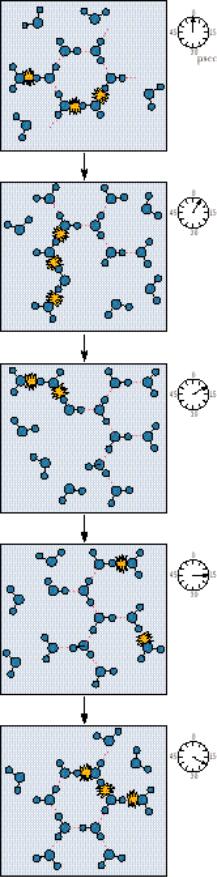
The present interpretation of water structure is that water molecules are connected by uninterrupted H bond paths running in every direction, spanning the whole sample. The participation of each water molecule in an average state of H bonding to its neighbors means that each molecule is connected to every other in a fluid network of H bonds. The average lifetime of an H-bonded connection between two H2O molecules in water is 9.5 psec (picoseconds, where 1 psec 10 12 sec). Thus, about every 10 psec, the average H2O molecule moves, reorients, and interacts with new neighbors, as illustrated in Figure 2.3.
In summary, pure liquid water consists of H2O molecules held in a random, three-dimensional network that has a local preference for tetrahedral geometry but contains a large number of strained or broken hydrogen bonds. The presence of strain creates a kinetic situation in which H2O molecules can switch H-bond allegiances; fluidity ensues.
Solvent Properties
Because of its highly polar nature, water is an excellent solvent for ionic substances such as salts; nonionic but polar substances such as sugars, simple alcohols, and amines; and carbonyl-containing molecules such as aldehydes and ketones. Although the electrostatic attractions between the positive and negative ions in the crystal lattice of a salt are very strong, water readily dissolves salts. For example, sodium chloride is dissolved because dipolar water molecules participate in strong electrostatic interactions with the Na and Cl ions, leading to the formation of hydration shells surrounding these ions (Figure 2.4). Although hydration shells are stable structures, they are also dynamic. Each water molecule in the inner hydration shell around a Na ion is replaced on average every 2 to 4 nsec (nanoseconds, where 1 nsec 10 9 sec) by another H2O. Consequently, a water molecule is trapped only several hundred times longer by the electrostatic force field of an ion than it is by the H-bonded network of water. (Recall that the average lifetime of H bonds between water molecules is about 10 psec.)
Water Has a High Dielectric Constant
The attractions between the water molecules interacting with, or hydrating, ions are much greater than the tendency of oppositely charged ions to attract one another. The ability of water to surround ions in dipole interactions and
● The fluid network of H bonds linking water molecules in the liquid state. It is revealing to note that, in 10 psec, a photon of light (which travels at 3 108 m/sec) would move a distance of only 0.003 m.
37

38 Chapter 2 ● Water, pH, and Ionic Equilibria
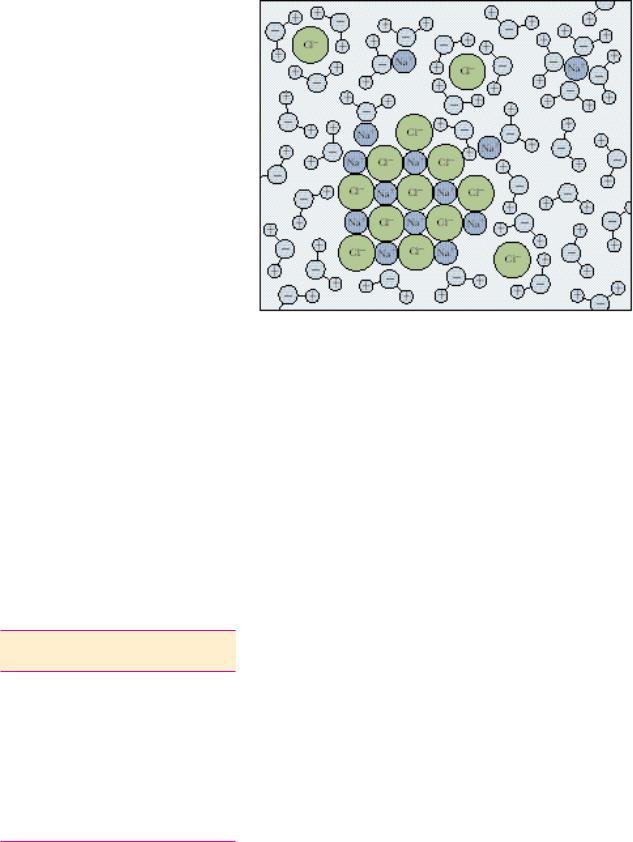
FIGURE 2.4 ● Hydration shells surrounding ions in solution. Water molecules orient so that the electrical charge on the ion is sequestered by the water dipole. For positive ions (cations), the partially negative oxygen atom of H2O is toward the ion in solution. Negatively charged ions (anions) attract the partially positive hydrogen atoms of water in creating their hydration shells.
diminish their attraction for one another is a measure of its dielectric constant, D. Indeed, ionization in solution depends on the dielectric constant of the solvent; otherwise the strongly attracted positive and negative ions would unite to form neutral molecules. The strength of the dielectric constant is related to the force, F, experienced between two ions of opposite charge separated by a distance, r, as given in the relationship
F e1e2/Dr2
Table 2.1
Dielectric Constants* of Some
Common Solvents at 25°C
Solvent |
Dielectric Constant (D) |
|
|
Water |
78.5 |
Methyl alcohol |
32.6 |
Ethyl alcohol |
24.3 |
Acetone |
20.7 |
Acetic acid |
6.2 |
Chloroform |
5.0 |
Benzene |
2.3 |
Hexane |
1.9 |
*The dielectric constant is also referred to as relative permittivity by physical chemists.
where e1 and e2 are the charges on the two ions. Table 2.1 lists the dielectric constants of some common liquids. Note that the dielectric constant for water is more than twice that of methanol and more than 40 times that of hexane.
Water Forms H Bonds with Polar Solutes
In the case of nonionic but polar compounds such as sugars, the excellent solvent properties of water stem from its ability to readily form hydrogen bonds with the polar functional groups on these compounds, such as hydroxyls, amines, and carbonyls. These polar interactions between solvent and solute are stronger than the intermolecular attractions between solute molecules caused by van der Waals forces and weaker hydrogen bonding. Thus, the solute molecules readily dissolve in water.
Hydrophobic Interactions
The behavior of water toward nonpolar solutes is different from the interactions just discussed. Nonpolar solutes (or nonpolar functional groups on biological macromolecules) do not readily H bond to H2O, and, as a result, such compounds tend to be only sparingly soluble in water. The process of dissolving such substances is accompanied by significant reorganization of the water surrounding the solute so that the response of the solvent water to such solutes can be equated to “structure making.” Because nonpolar solutes must occupy space, the random H-bond network of water must reorganize to accommodate them. At the same time, the water molecules participate in as many H-bonded
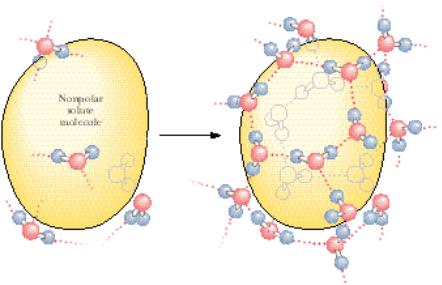
interactions with one another as the temperature permits. Consequently, the H-bonded water network rearranges toward formation of a local cagelike (clathrate) structure surrounding each solute molecule (Figure 2.5). This fixed orientation of water molecules around a hydrophobic “solute” molecule results in a hydration shell. A major consequence of this rearrangement is that the molecules of H2O participating in the cage layer have markedly reduced orientational options. Water molecules tend to straddle the nonpolar solute such that two or three tetrahedral directions (H-bonding vectors) are tangential to the space occupied by the inert solute. This “straddling” means that no water H-bonding capacity is lost because no H-bond donor or acceptor of the H2O is directed toward the caged solute. The water molecules forming these clathrates are involved in highly ordered structures. That is, clathrate formation is accompanied by significant ordering of structure or negative entropy.
Under these conditions, nonpolar solute molecules experience a net attraction for one another that is called hydrophobic interaction. The basis of this interaction is that when two nonpolar molecules meet, their joint solvation cage involves less surface area and less overall ordering of the water molecules than in their separate cages. The “attraction” between nonpolar solutes is an entropy-driven process due to a net decrease in order among the H2O molecules. To be specific, hydrophobic interactions between nonpolar molecules are maintained not so much by direct interactions between the inert solutes themselves as by the increase in entropy when the water cages coalesce and reorganize. Because interactions between nonpolar solute molecules and the water surrounding them are of uncertain stoichiometry and do not share the equality of atom-to-atom participation implicit in chemical bonding, the term hydrophobic interaction is more correct than the misleading expression hydrophobic bond.
Amphiphilic Molecules
Compounds containing both strongly polar and strongly nonpolar groups are called amphiphilic molecules (from the Greek amphi meaning “both,” and philos meaning “loving”), also referred to as amphipathic molecules (from the Greek pathos meaning “passion”). Salts of fatty acids are a typical example that
2.1 ● Properties of Water |
39 |
FIGURE 2.5 ● Formation of a clathrate structure by water molecules surrounding a hydrophobic solute.
amphiphilic molecules, amphipathic molecules ● compounds containing both strongly polar and strongly nonpolar groups
40 Chapter 2 ● Water, pH, and Ionic Equilibria
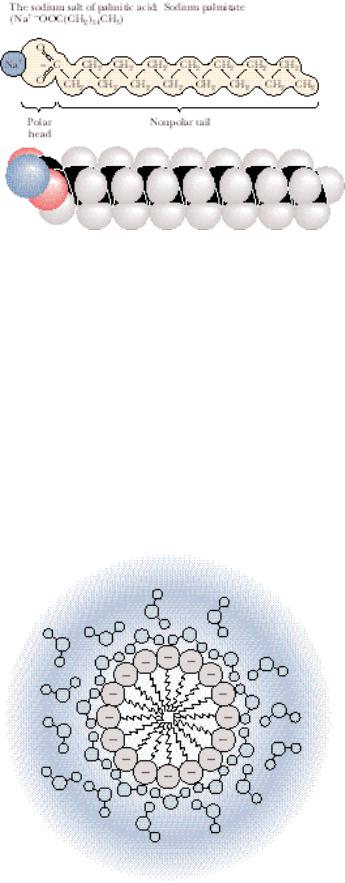
FIGURE 2.6 ● An amphiphilic molecule: sodium palmitate. Amphiphilic molecules are frequently symbolized by a ball and zig-zag line structure,  , where the ball represents the hydrophilic polar head and the zig-zag represents the nonpolar hydrophobic hydrocarbon tail.
, where the ball represents the hydrophilic polar head and the zig-zag represents the nonpolar hydrophobic hydrocarbon tail.
has biological relevance. They have a long nonpolar hydrocarbon tail and a strongly polar carboxyl head group, as in the sodium salt of palmitic acid (Figure 2.6). Their behavior in aqueous solution reflects the combination of the contrasting polar and nonpolar nature of these substances. The ionic carboxylate function hydrates readily, whereas the long hydrophobic tail is intrinsically insoluble. Nevertheless, sodium palmitate and other amphiphilic molecules readily disperse in water because the hydrocarbon tails of these substances are joined together in hydrophobic interactions as their polar carboxylate functions are hydrated in typical hydrophilic fashion. Such clusters of amphipathic molecules are termed micelles; Figure 2.7 depicts their structure. Of enormous biological significance is the contrasting solute behavior of the two ends of amphipathic molecules upon introduction into aqueous solutions. The polar ends express their hydrophilicity in ionic interactions with the solvent, whereas their nonpolar counterparts are excluded from the water into a hydrophobic domain constituted from the hydrocarbon tails of many like molecules. It is this behavior that accounts for the formation of membranes, the structures that define the limits and compartments of cells (see Chapter 9).
FIGURE 2.7 ● Micelle formation by amphiphilic molecules in aqueous solution. Negatively charged carboxylate head groups orient to the micelle surface and interact with the polar H2O molecules via H bonding. The nonpolar hydrocarbon tails cluster in the interior of the spherical micelle, driven by hydrophobic exclusion from the solvent and the formation of favorable van der Waals interactions. Because of their negatively charged surfaces, neighboring micelles repel one another and thereby maintain a relative stability in solution.
